Registration is often needed to quantify breast cancer change over time. This is especially important to monitor the change of breast cancer patient, and access their response to the chemotherapy (treatment effects). It is also one of the first steps towards differentiation between complete responders (subjects who show the absence of any residual invasive cancer in the breast and the absence of any metastatic cells in the regional lymph nodes) and non-complete-responders.
Below is an example. DRAMMS recovers the deformation from post to pre chemotherapy in this breast cancer patient.
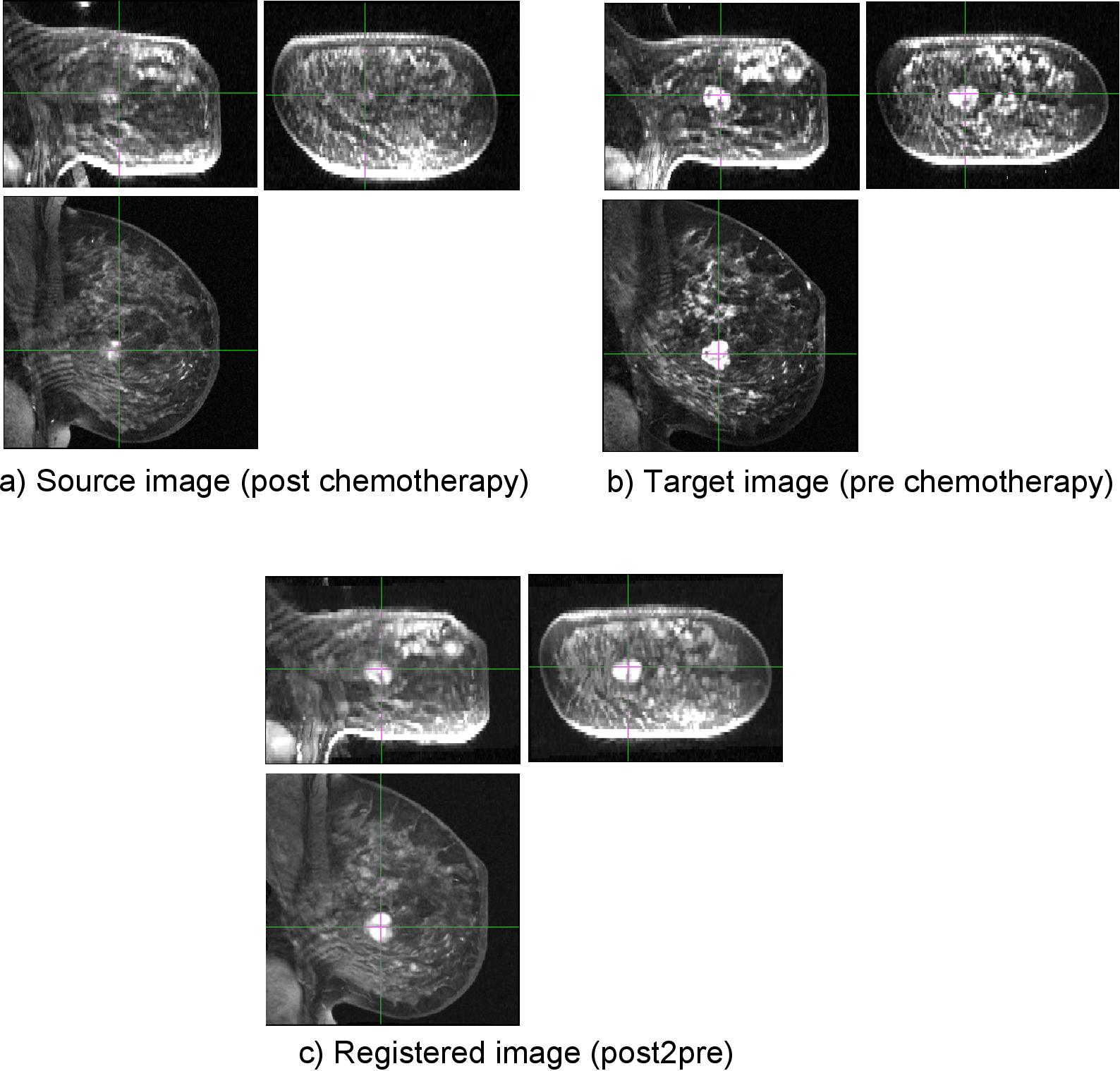
Registration of breast cancer images for the same subject, to monitor the effect of chemotherapy in altering breast cancer over time.
dramms -S src_breastPost.hdr -T trg_breastPre.hdr
-O src2trg.nii.gz -D def_S2T.nii.gz -g 0.3
Registering this pair of 3D images (target image 256 x 256 x 64 voxels, 0.78 x 0.78 x 2.30 mm^3/voxel) takes 9.1 GB memory and finishes in 81 minutes in Linux OS (2.80GHz CPU).
If one can afford less memory, please use -u option to choose memory usage in different levels (the lowest being about 1/4 of maximum memory used). This may however slightly reduce registration accuracy.
With the DRAMMS registration, we can recover the deformation/change over time at the voxel level. This is done by the Jacobian Determinant map calculated from the DRAMMS deformation (click here for how to calculate Jacobian Determinant maps). Below is an example.
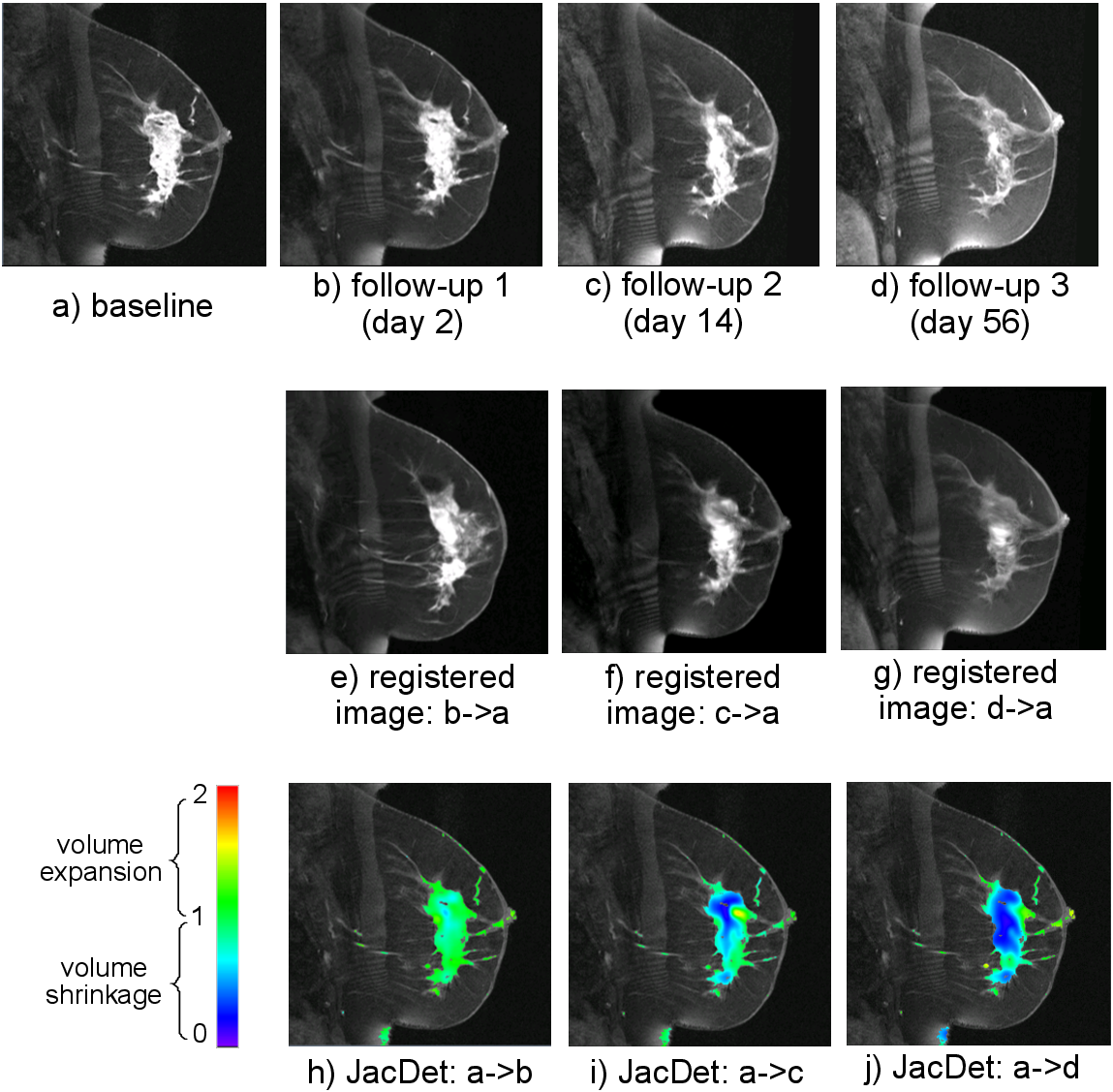
Quantification of the longitudinal cancer of a breast cancer patient, by the Jacobian Determinant maps derived from the DRAMMS-computed deformation.